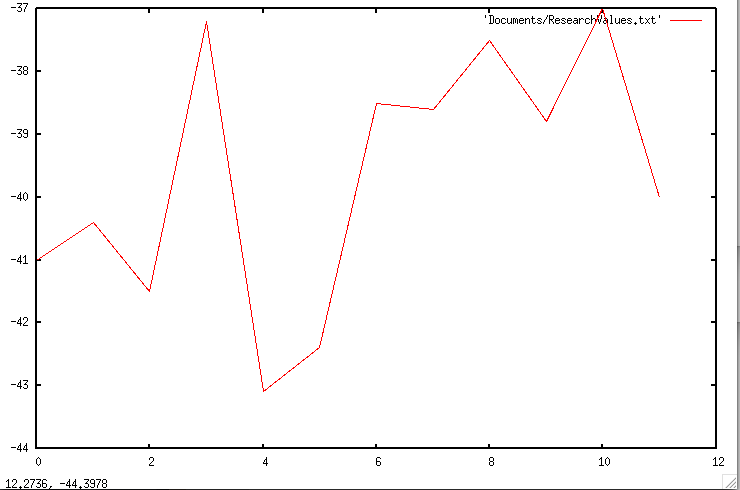
This week I have adapted my program to run on the Ubuntu lab computer. I configured and installed Mfold, Rfold, and UNAFold to allow my program to perform system calls. Now, instead of using an rfold file that already exists in a directory, my program can make calls directly to rfold (several times) and generate the necessary files itself (for a short synopsis of my program subroutines, see below). Later today, I will upload additional graphs of the data my program has generated.
importGlobalSequence -> Imports the sequence
calcFoldingEnergy -> Calculates Folding Energy using Mfold
importLocalSequence -> Selects a K-mer
generateRfoldFile -> Generates Rfold Probability File
createArrOfPosAboveThreshold -> Creates array of prob. above threshol
modifySequence -> Modifies the sequence
generateModifiedFastaFile-> Saves the modified Fasta Sequence
synonymousChange-> Creates a synonymous change
importGlobalSequence -> Imports the sequence
calcFoldingEnergy -> Calculates Folding Energy using Mfold
importLocalSequence -> Selects a K-mer
generateRfoldFile -> Generates Rfold Probability File
createArrOfPosAboveThreshold -> Creates array of prob. above threshol
modifySequence -> Modifies the sequence
generateModifiedFastaFile-> Saves the modified Fasta Sequence
synonymousChange-> Creates a synonymous change
 RSS Feed
RSS Feed
